Introduction
I just joined a competition called ‘Three Minute Thesis’ at my university, in which I have to explain, in three minutes, what my thesis is about*. The thing is that my thesis is about detecting exoplanet (that is, planets outside our solar system) atmospheres from the ground through an effect called transmission spectroscopy but, although that sounds cool for most people (except when I mention ‘transmission spectroscopy’, which sounds like a magical, frightening and strange word for non-astronomers), I don’t really know if they understand how hard this stuff is to do. In order to show this, I want to share a little order-of-magnitude calculation that I did in order to show how hard it is to find exoplanet atmospheres through this method.
What is transmission spectroscopy?
As of today, most exoplanets discovered so far have been discovered through the method of transits: the apparent decrease in flux of a star due to a planet passing in front of it. This is, of course, a lucky event: for earth-like planets (that is, planets orbiting Sun-like stars at the same distance as the Earth), assuming orbits around stars are totally random, the probability of finding a transiting planet is ! When we are lucky enough to see this event, we can exploit a lot of information concerning the atmosphere of the planet. The effect I’m going to talk about in this post comes from a very simple idea: during a transit, part of the light gets blocked by the planet passing in front of the star but some of it also goes through the atmosphere, so the informatlon regarding the atmosphere gets imprinted on the stellar light that we receive at Earth. This is why the effect is termed transmission spectroscopy; stellar light gets transmitted through the atmosphere of the planet, and we try to detect that signature. A picture is worth more than a thousand words, so here’s a diagram I made:
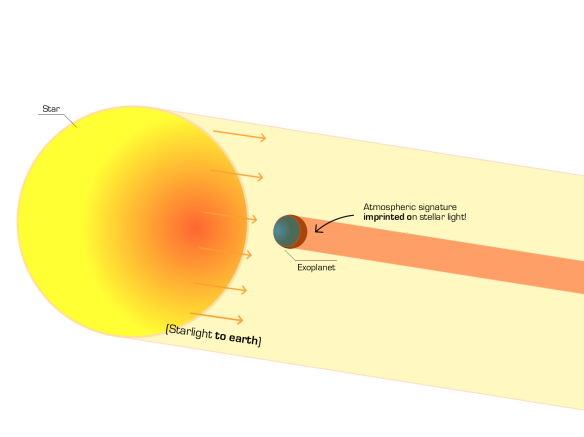
Diagram showing how light is not only blocked by a transiting planet during a transit event, but also how the atmospheric signature (red ring around the blue planet) is imprinted on the stellar light. Credits: Néstor Espinoza.
Seems like a cool effect, huh? It also seems like a very small effect. Let’s first explore how small of an effect a transiting planet makes and, then, jump into the very small effect the atmosphere might be able to produce!
The tiny effect of transits
Let’s consider a planet of radius transiting a star of radius
. In the absence of an atmosphere, during a transit, a planet would imprint a decrease in the observed stellar flux (energy per unit time per unit area) at first order (and assuming an orbit very close to the star) equal to the ratio of the observed areas, i.e.,
. How small is this effect? For the case of a Jupiter-sized planet transiting a star like the Sun (where the ratio of radii is
),
, or 1% of the stellar flux! To have an idea of how small this is, let’s first try to understand how dim a star is as seen from Earth. Consider a star like the sun at 4 lightyears away (the distance to the Alpha Centauri system for which, to date, no transiting planet has been detected) with a transiting planet like Jupiter. If we consider its luminosity to be solar, i.e.
, the observed flux at Earth of this star would be
.
To compare this flux, let’s consider a lightbulb, which at best has an output of around 100 W. In order to observe a similar flux as that of the star mentioned before, you would need to be at a distance
from it. For the ones out there that have come to Santiago de Chile, this distance is roughly the distance from the top of San Cristobal hill to the farthest lightbulb you can see. This photo might enlighten your imagination:
Pretty dim, huh? Now, how small is the effect of a 1% decrease in flux due to a transiting planet? Well, remember that a 1% decrease in flux for a star’s flux happens when a planet 1/10 of its radius passes in front of it when the star is like our Sun. The farthest lightbulb you can see in the photo comes from a source with a radius not bigger than, say, ~10 cm; a 1% decrease in flux in that lightbub then can arise from an object 1/10 of its size passing in front of it, or, an object with a radius of ~1 cm! That’s like detecting the effect of a moth passing between you and that lightbulb very close to it! That is a really small effect. However, as I mentioned in the introduction, most planets have been detected in this way, and there are large collaborations of astronomers trying to detect more exoplanets like these ones (have you ever heard of HATSouth, for example?). We might say that this is one of the most successful exoplanet finding tools to date! Now, If you think the effect of transits is small, wait until you see the really, really small effect that the atmosphere imprints on exoplanetary transits.
The (really) tiny effect of transmission spectroscopy
Let’s consider the same planet as before, but let’s now assume that it has an atmosphere. The effect that one usually wants to detect is the absorption feature of some atomic and/or molecular feature and, therefore, what one usually seeks is a transit event observed in different wavelengths. If there is absorption in a given wavelength, then the planet will look bigger than in other wavelengths. To simplify the matters, let’s assume that the whole atmosphere is of this given element that absorbs all the light at a given wavelength. If we assume that the only force that keeps this element from escaping to outer space is gravity, then it should be a given height where gravity is so small that the element can escape due to the internal kinetic energy of the compound. If the kinetic energy is of the same order of magnitude as the gravitational potential, then the gas will start escaping from the atmosphere. Assuming the element behaves more or less like an ideal gas and is more or less isothermal, the mean kinetic energy of it is given by
where is Boltzmann’s constant and
is the temperature of the gas in Kelvins. On the other hand, if the element has a mass
, the graviational potential is given by
where is the gravitational acceleration of the planet and
is the height at which this potential is measured. Now, when
, the kinetic energy is enough to let some of the element escape to outer space. This occurs at a height
This number is usually known as the scale height, and it defines a characteristic height to a given atmosphere (its actually the height at which an atmosphere in thermodynamic and hydrostatic equilibrium decreases its presure by a factor ). For Earth, for example,
,
K and the mean mass of the constituents is
, where
is the mass of a proton. This gives
which is the characteristic height of our atmosphere. Of course, this is not the height of the whole atmosphere that would be probed by the effect of transmission spectroscopy; this largely depends on the constituents, layers and properties of those layers in each atmosphere (i.e., the pressure-temperature profile). As a good average between thin () and very extended (
) atmospheres,
seem to be a reasonable choice. Let’s now assume then that our close-in Jupiter transiting planet has an atmosphere with a height of
. Most of the close-in Jupiters have been discovered at periods around
days, which implies temperatures close to
K, reason by which these planets are usually called “hot” Jupiters. If we assume it to be an almost exact Jupiter analog, its gravity should be
. Furthermore, it should be made mainly of hydrogen gas, just as Jupiter, so
. Combining all of that information, the scale-height for our Hot Jupiter should be
In this case then, if the atmosphere were to absorb all the light at a given wavelength, the planet would appear to have a radius of . This induces a transit signal equal to the ratios of the areas, but now with the “new” planet + atmosphere radius,
or, in a more compact form,
where we have ommited the term (its very small compared to the other terms), used the term
already shown before and defined the term
. Note that is this last term that is added to the “normal” transit signal (
), so we can interpret it also as a percentage change in flux due solely to the atmosphere. Using the radius of Jupiter to be
, a radius of the Sun of
and the calculated scale-height, we obtain,
or 0.03%! That’s two orders of magnitude smaller than the transit signal which was 1%! If we go back to our example of the lightbulb again, recalling that a change in flux of
can arise from the ratio of the areas of the two objects, if our lightbulb has a
radius, then a change in flux of
can come from an object of size
passing close-by, in front and between us and the lightbulb. That’s measuring the effect of a 2 mm object passing in front of the lightbulb! Enough precision to give the moth a coat for the winter or a bathing suit for the summer! Despite the effect being very small, this effect has been measured successfully both from space and ground based observatories. However, these studies have been done only for a handful of systems; there are many more problems than just getting enough light from the stars in order to detect this effect, where ground based observations are by far the most challenging ones. Imagine trying to measure that 0.03% flux change from the lightbulb, but with fog, while your instrument moves, rotates and shakes due to its own gravity…those systematic effects are the biggest problems in current ground-based measurements. It’s quite a challenge, and that’s actually what makes it so interesting! Frontier science usually comes from beating those challenges with clever ideas so, stay tuned!
Conclusions
Detecting exoplanet atmospheres is hard. Very hard. In the best-case scenario, It is analogous to trying to measure the flux change in a lightbulb 20 km away from your location with enough precision to build a coat or a swimsuit for a moth passing in front and very close to the lightbulb. The signal depends largely on temperature, gravity and the mass of the elements in the atmosphere; it favours higher temperatures, lower gravities and lighter constituents. The biggest challenge, though, is not the signal but the actual systematic effects the instruments and the Earth’s atmosphere (for the case of ground-based observations) imprints on the signals but various groups of astronomers are working on this as we speak…stay tuned for future discoveries!
*: As an update on this, I won the People’s Choice award in the competition! It was really cool, specially because there was no people from my department in the audience (who were the ones who voted for me).